Notre Equipe
Your content goes here. Edit or remove this text inline or in the module Content settings. You can also style every aspect of this content in the module Design settings and even apply custom CSS to this text in the module Advanced settings.











Karima EL MAHBOUBI















Anciens membres
Charles Uhlmann @UhlmannCharles (Master 2 student, now Ph.D student in J. Parker’s lab, Cologne, Germany)
Ahmed Khamassi (Master 2 student, now Ph.D student at INSA, Toulouse)
Chloé Langlet (« Licence pro » student, now Master student at UPS)
Nicolas Vigneron (PhD student, now postdoc HES-SO University, Switzerland)
Roland Berdaguer (Master 2 student, now Ph. D student at Wageningen University, NL)
Gaetan Tressière (Master 2 student)
Léa Boyrie (PhD student)
Simon Pons (PhD student)
Duchesse Mbadinga (PhD student)
Thèmes de Recherche
Evolution of Plant – Microbe Interactions
Plants interact with a huge diversity of microorganisms in associations ranging from parasitic to mutualistic, with either bacteria or filamentous fungi or oomycetes.
Since plants colonized lands 450 million years ago, these associations have evolved in opposite directions. Mutualism has been maintained for hundreds of million years whereas the interactions with parasites are fast evolving.
Studying the interactions between plant lineages spanning 500 milion years of diversification and multiple microorganisms, our goals are to discover the plant molecular mechanisms behind mutualistic and parasitic interactions with microorganisms and to understand how they evolved.
MUTUALISTIC SYMBIOSES
Mutualism between plants and microorganisms has been essential for the evolution of terrestrial ecosystems for millions of years. It has been proposed that even the colonization of lands by plants was facilitated by a mutualistic symbiosis formed with arbuscular mycorrhizal fungi. This symbiosis, by far the most widespread in land plants, results in the accommodation of the symbiotic fungus inside the plant cells. Following this initial symbiosis, multiple other intracellular symbioses have evolved in plants as diverse as orchids, Ericaceae such as cranberry, legumes or the Jungermanniales, a group of bryophytes. These symbioses provide numerous benefits, improving plant nutrient acquisition and fitness. Despite their absolute importance in terrestrial ecosystems, the molecular mechanisms underlying the origin and subsequent evolution of intracellular symbioses in plants remain poorly understood.
Our research aims at understanding how these symbiotic associations evolved. Using combinations of phylogenomics, biochemistry and genetics in multiple plants we try to determine the molecular bases of the origin, conservation and diversification of these associations.

PLANT – PATHOGEN INTERACTIONS
In their constantly changing environment, plants face a large diversity of potentially pathogenic microorganisms.
The ability to limit the infection by these parasites varies from species to species and, even, between accessions of a given species.
The molecular mechanisms mediating resistance have been widely studied in flowering plants.
By contrast, how non-vascular plants such as liverworts defend themselves from pathogens and the genetics behind remains poorly characterized.
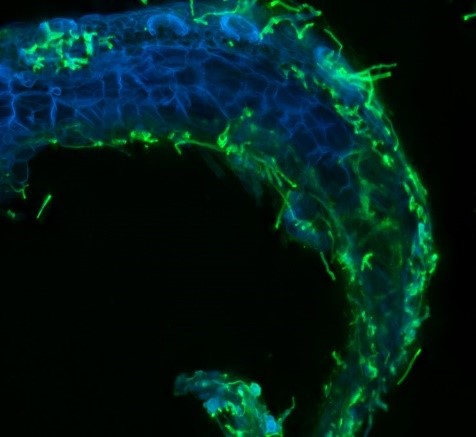

INTERPLAY BETWEEN SYMBIOSIS AND IMMUNITY
Interactions between organisms affect gene and ecosystem diversity. Given that most species are constantly challenged by both mutualistic and parasitic microorganisms, we propose that mutualism and parasitism could influence each other. Here, using plants as a model we will combine our complementary expertise to test the hypothesis that parasitism and mutualism impact the evolution of each other. To do this, we first propose to identify genes involved in parasitism or mutualism using genetic approaches (GWAS, selection scans) and to compare their evolution within species of two deeply divergent clades of land plants, angiosperms and liverworts, including in species which have lost mutualism (e.g. Arabidopsis thaliana and Marchantia polymorpha). Then, through phylogenomic approaches across the entire embryophyte phylogeny, we will finely describe the evolution of these genes and infer selective processes to understand how selection can manage the interplay between mutualism and parasitism.
FUNDINGS
Publications
- Hernandez-Reyes C, Lichtenberg E, Keller J, Delaux P-M, Ott T, Schenk ST. (2021) NIN-Like Proteins interesting Players in Rhizobia-Induced Nitrate Signaling Response during Interaction with Non-Legume Host Arabidopsis thaliana. MPMI, 10.1094/MPMI-10-21-0261-R
- Vásquez-Ocmína PG, Marti G, Bonhomme M, Mathis F, Fournier S, Bertani S, Maciuk A. (2021) Cannabinoids vs. whole metabolome: Relevance of cannabinomics in analyzing Cannabis varieties. Analytica Chimica Acta, 10.1016/j.aca.2021.339020
- Kodama K*, Rich MK*, Yoda A, Shimazaki S, Xie X, Akiyama K, Mizuno Y, Komatsu A, Luo Y, Suzuki H, Kameoka H, Libourel C, Keller J, Sakakibara K, Nishiyama T, Nakagawa T, Mashiguchi K, Uchida K, Yoneyama K, Tanaka Y, Yamaguchi S, Shimamura M, Delaux P-M#, Nomura T#, Kyozuka J#. (2021) An ancestral function of strigolactones as symbiotic rhizosphere signals. bioRxiv 10.1101/2021.08.20.457034
- El Mahboubi K & Delaux P-M. (2021) Plant biology: Two green revolutions mediated by DELLA. Current Biology, 10.1016/j.cub.2021.07.024
- Rich MK*, Vigneron N*, Libourel C, Keller J, Xue L, Hajheidhari M, Radhakrishnan GV, Le Ru A, Diop IS, Potente G, Conti E, Duijsings D, Batut A, Le Faouder P, Kodama K, Kyozuka J, Sallet E, Bécard G, Rodriguez-Franco M, Ott T, Bertrand-Michel J, Oldroyd GED, Szövényi P, Bucher M, Delaux P-M. (2021) Lipid exchanges drove the evolution of mutualism during plant terrestrialization. Science, 10.1126/science.abg0929 Download the .pdf : science.sciencemag.org/cgi/content/full/372/6544/864?ijkey=GNoVgjh4AJNZE&keytype=ref&siteid=sci
- Liang P, Schmitz C, Lace B, Ditengou FA, Keller J, Libourel C, Delaux P-M, Ott T. (2021) A formin-mediated cell wall- plasma membrane- cytoskeleton continuum is required for symbiotic infections in Medicago truncatula. bioRxiv, 10.1101/2020.07.10.197160 | Current Biology, 10.1016/j.cub.2021.04.002
- Bonhomme M, Bensmihen S, André O, Amblard E, Garcia M, Maillet F, Puech-Pagès V, Gough C, Fort S, Cottaz S, Bécard G, Jacquet C. (2021) Distinct genetic basis for root responses to lipo-chitooligosaccharide signal molecules from distinct microbial origins. biorXiv, 10.1101/2020.09.09.285668 | J Exp Bot, 10.1093/jxb/erab096
- Delaux P-M* & Schornack S* (2021) Plant evolution driven by interactions with symbiotic and pathogenic microbes. Science, 10.1126/science.aba6605
- Mathu M*, Kruger M*, Kruger C*, Wang Y, Stajich JE, Keller J, Chen ECH, Yildirir G, Villeneuve-Laroche M, Roux C, Delaux P-M, Corradi N. (2021) The genome of Geosiphon pyriformis reveals ancestral traits linked to the emergence of the arbuscular mycorrhizal symbiosis. Current Biology, 10.1016/j.cub.2021.01.058
- Isidra-Arellano MC, Delaux P-M, Valdés-López O. (2021) The Phosphate Starvation Response System: its role in the regulation of plant-microbe interactions. Plant Cell Physiol, 10.1093/pcp/pcab016
- Quilbé J, Lamy L, Brottier L, Leleux P, Fardoux J, Rivallan R, Benichou T, Guyonnet R, Becana M, Villar I, Garsmeur O, Hufnagel B, Delteil A, Gully D, Chaintreuil C, Pervent M, Cartieaux F, Bourge M, Valentin N, Martin G, Fontaine L, Droc G, Dereeper A, Farmer A, Libourel C, Nouwen N, Gressent F, Mournet P, D’Hont A, Giraud E, Klopp C, Arrighi JF. (2021) Genetics of nodulation in Aeschynomene evenia uncovers mechanisms of the rhizobium-legume symbiosis. Nature Communications, 10.1038/s41467-021-21094-7.
- Pons S, Fournier S, Chervin C, Bécard G, Rochange S, Frei Dit Frey N, Puech Pagès V. Phytohormone production by the arbuscular mycorrhizal fungus Rhizophagus irregularis. PLoS One, 10.1371/journal.pone.0240886
- Aoun N, Desaint H, Boyrie L, Bonhomme M, Deslandes L, Berthomé R, Roux F (2020). A complex network of additive and epistatic quantitative trait loci underlies natural variation of Arabidopsis thaliana quantitative disease resistance to Ralstonia solanacearum under heat stress. Molecular Plant Pathology, doi.org/10.1111/mpp.12964
- Bonhomme M*, Bensmihen S*, Andre O, Amblard E, Garcia M, Maillet F, Puech-Pages V, Gough C, Fort S, Cottaz S, Becard G, Jacquet C. (2020) Distinct genetic bases for plant root responses to lipo-chitooligosaccharide signal molecules from distinct microbial origins. biorXiv, 10.1101/2020.09.09.285668
- Frei-dit-Frey N & Favery B. (2020) Plant‐parasitic nematode secreted peptides hijack a plant secretory pathway. New Phytologist, 10.1111/nph.16842
- Boyrie L, Moreau C, Frugier F, Jacquet C, Bonhomme M. (2020) A linkage disequilibrium-based statistical test for Genome-Wide Epistatic Selection Scans in structured populations. Heredity, 10.1038/s41437-020-0349-1 | biorXiv: 10.1101/2020.02.14.949206
- Liang P, Schmitz C, Lace B, Ditengou FA, Keller J, Libourel C, Delaux P-M, Ott T. (2020) A formin-mediated cell wall- plasma membrane- cytoskeleton continuum is required for symbiotic infections in Medicago truncatula. bioRxiv, 10.1101/2020.07.10.197160
- Rush TA, Puech-Pagès V, Bascaules A, Jargeat P, Maillet F, Haouy A, Maës AQ, Carrera Carriel C, Khokhani D, Keller-Pearson M, Tannous J, Cope KR, Garcia K, Maeda J, Johnson C, Kleven B, Choudhury QJ, Labbé J, Swift C, O’Malley MA, Bok JW, Cottaz S, Fort S, Poinsot V, Sussman MR, Lefort C, Nett J, Keller NP, Bécard G, Ané JM (2020) Lipo-chitooligosaccharides as regulatory signals of fungal growth and development. Nature Communications, 10.1038/s41467-020-17615-5
- Berbee ML, Strullu-Derrien C, Delaux P-M, Strother PK, Kenrick P, Selosse M-A, Taylor JW. (2020) Genomic and fossil windows into the secret lives of the most ancient fungi. Nature Review Microbiology, 10.1038/s41579-020-0426-8
- Fraisier-Vannier O, Chervin J, Cabanac G, Puech-Pages V, Fournier S, Durand V, Amiel A, Andre O, Abdelaziz Benamar O, Tsugawa H, Dumas B, Marti G. (2020) MS-CleanR: A feature-filtering workflow for untargeted LC-MS based metabolomics. Analytical Chemistry, 10.1021/acs.analchem.0c01594
- Rich MK & Delaux P-M (2020) Plant Evolution: When Arabidopsis is more ancestral than Marchantia. Current Biology, 10.1016/j.cub.2020.04.077
- Taulera Q, Lauressergues D, Martin K, Cadoret M, Servajean V, Boyer F-D, Rochange S. (2020) Initiation of arbuscular mycorrhizal symbiosis involves a novel pathway independent from hyphal branching. Mycorrhiza, 10.1007/s00572-020-00965-9
- Rathgeb U, Chen M, Buron F, Feddermann N, Schorderet M, Raisin A, Marc-Martin S, Haeberli G-Y, Keller J, Delaux P-M, Schaefer D, Reinardt D. (2020) VAPYRIN-like is required for development of the moss Physcomitrella patens. Development, 10.1242/dev.184762
- Keller J & Delaux P-M. (2020) Evolution of plant metabolism: A (bio)synthesis. Current Biology, 10.1016/j.cub.2020.03.020
- Su C, Klein M-L, Hernandez-Reyes C, Batzenschlager M, Ditengou FA, Keller J, Delaux P-M, Ott T. (2020) The Medicago truncatula DREPP protein triggers microtubule fragmentation in membrane nanodomains during symbiotic infections. Plant Cell, doi.org/10.1105/tpc.19.00777 | biorXiv: doi.org/10.1101/794008
- Li F-W, Nishiyama T, Waller M, Frangedakis E, Keller J, Li Z, Fernandez-Pozo N, Barker MS, Bennett T, Blázquez MA, Cheng S, Cuming AC, de Vries J, de Vries s, Delaux P-M, Diop IS, Harrison J, Hauser D, Hernández-García J, Kirbis A, Meeks JC, Monte I, Mutte SK, Heubauer A, Quandt D, Robison T, Shimamura M, Rensing SA, Villarreal JC, Weijers D, Wicke s, Wong GKS, Sakakibara K, Szövényi P. (2020) Anthoceros genomes illuminate the origin of land plants and the unique biology of hornworts. Nature Plants, 10.1038/s41477-020-0618-2
- Radhakrishnan GV*, Keller J*, Rich MK*, Vernié T*, Mbadinga Mbaginda DL, Vigneron N, Cottret L, San Clemente H, Libourel C, Cheema J, Linde AM, Eklund DE, Cheng S, Wong GKS, Lagercrantz U, Li F-W, Oldroyd GED, Delaux P-M. (2020) An ancestral signalling pathway is conserved in intracellular symbioses-forming plant lineages. Nature Plants, 10.1038/s41477-020-0613-7 | biorXiv: 10.1101/804591
- Hufnagel B, Marques A, Soriano A, Marquès L, Divol F, Doumas P, Sallet E, Mancinotti D, Carrere S, Marande W, Arribat S, Keller J, Huneau C, Blein T, Aime D, Laguerre M, Taylor J, Schubert V, Nelson M, Geu-Flores F, Crespi M, Gallardo-Guerrero K, Delaux P-M, Salse J, Bergès H, Guyot R, Gouzy J, Péret B. (2020) High-quality genome sequence of white lupin provides insight into soil exploration and seed quality. Nature Communications, 10.1038/s41467-019-14197-9
- Girardin A, Wang T, Ding Y, Keller J, Buendia L, Gaston M, Ribeyre C, Gasciolli V, Auriac C, Vernié T, Bendahmane A, Ried MK, Parniske M, Morel P, Vandenbussche M, Schorderet M, Reinhardt D, Delaux P-M, Bono J-J, Lefebvre B. (2019) LCO receptors involved in arbuscular mycorrhiza are functional for rhizobia perception in legumes. Current Biology, 10.1016/j.cub.2019.11.038
- Cheng S, Xian W, Fu Y, Marin B, Keller J, Wu T, Sun W, Li X, Xu Y, Zhang Y, Wittek S, Reder T, Günther G, Gontcharov A, Wang S, Li L, Liu X, Wang J, Yang H, Xu X, Delaux P-M, Melkonian B, Wong GKS, Melkonian M. (2019) Genomes of subaerial Zygnematophyceae provide insights into land plant evolution. Cell, 10.1016/j.cell.2019.10.019
- Delaux P-M#, Hetherington AJ, Coudert Y, Delwiche C, Dunand C, Gould S, Kenrick P, Li FW, Philippe H, Rensing SA, Rich M, Strullu-Derrien C, de Vries J (2019) Reconstructing trait evolution in plant evodevo studies. Current Biology, 10.1016/j.cub. 201909.04
Theses
Lea Boyrie, « Détection de gènes coadaptés par analyse pangénomique de signatures de sélection épistatique : application chez la légumineuse modèle Medicago truncatula », http://thesesups.ups-tlse.fr/4856/
Nicolas Vigneron, « Caractérisation des bases moléculaires de l’évolution de la symbiose mycorhizienne à arbuscules chez les plantes », http://thesesups.ups-tlse.fr/4594/
Duchesse Mbadinga, « Evolution de la cutine chez les plantes et son rôle durant la terrestrialisation »
umr5546@lrsv.ups-tlse.fr
Téléphone
Standard : 05.34.32.38.01
Fax : 05.34.32.38.02
Nous Trouver
24, chemin de Borde-Rouge.BP 42617 Auzeville.
31326, Castanet-Tolosan. FRANCE











